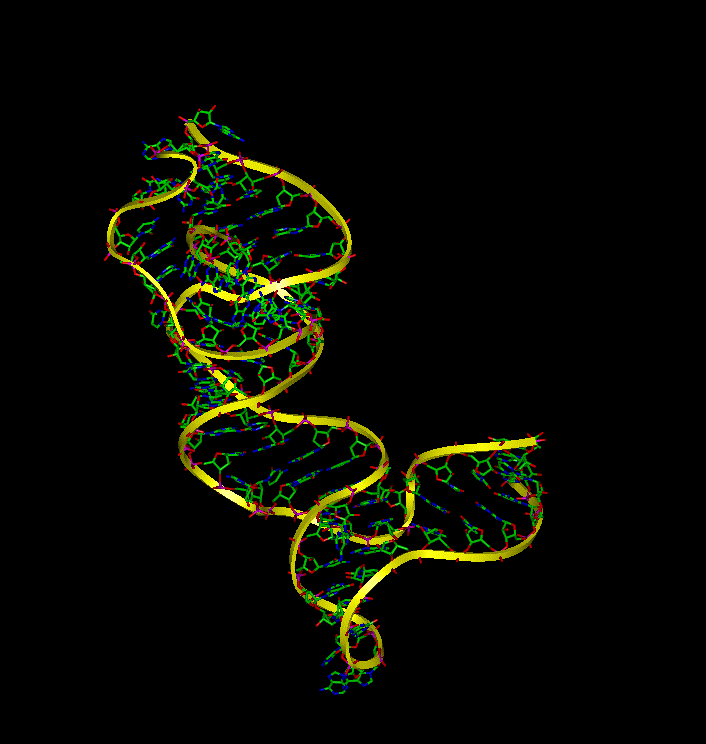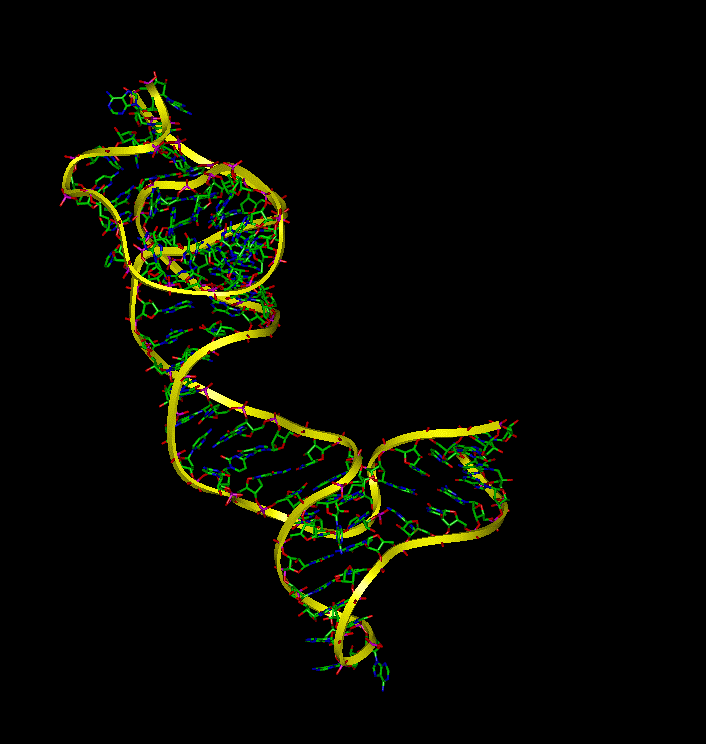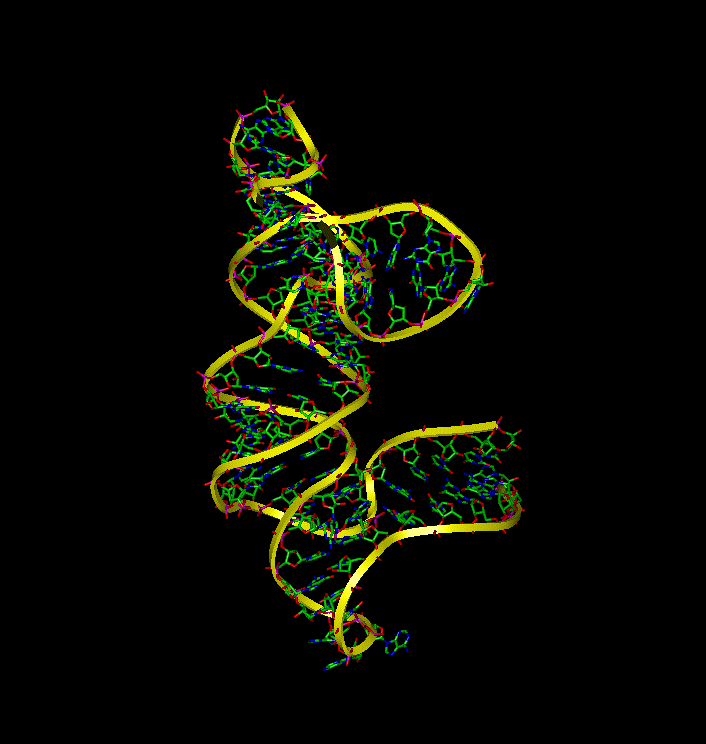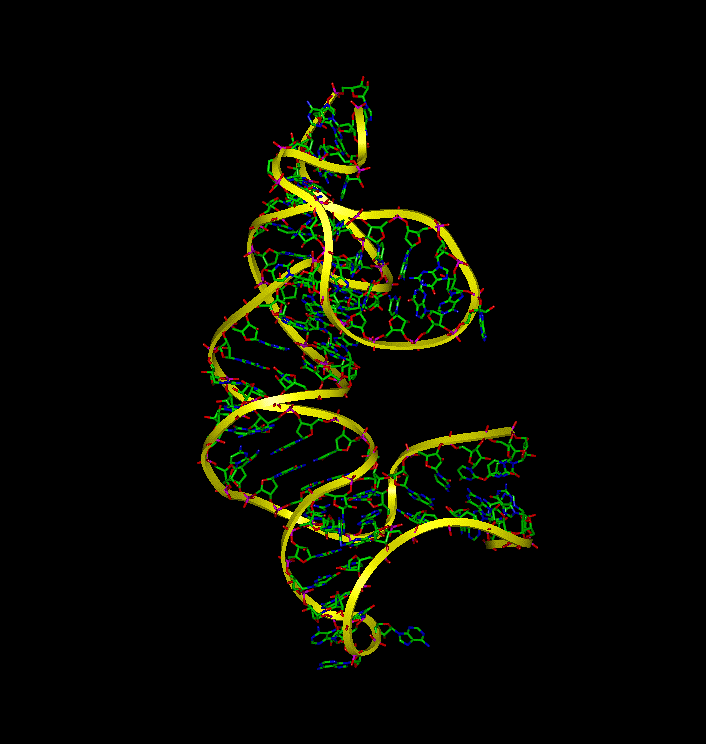
Molecular Dynamic simulations were utilized to determine the dynamic behavior of 16S rRNA in the presence and absense of S15 and to identify the binding interactions between the molecules. The simulations demonstrate that (a) 16S rRNA remains in a highly folded structure when bound to S15, (b) in the absence of S15, 16S rRNA alters its conformation and forms conformations similar to the bound that make it available for binding with S15, and (c) the unbound rRNA spends the majority of the time in an extentended conformation.
An equilibrium simulation was performed to determine the conformation of the rRNA-S15 complex. The simulation showed that the complex's conformation during equilibrium is similar to the its crystal structure. Throughout the equilibration, the rRNA remains coaxially stacked and the relative orientation of helix 20 and helix 22 remains similar to that found in the crystal structure.
To examine the dynamic behavior of 16S rRNA when S15 is removed from the complex, an additional equilibrium simulation was performed. Without the S15 protien, the 16S rRNA complex significantly fluctuates during equilibrium rather than remaining in the crystal structure. The image to the right depicts this fluctuation.